Clustering with Annealing#
This tutorial will cover clustering using PyQUBO and Openjij as an example for an application of annealing.
Clustering#
Assuming is given externally, we divide the given data into clusters. Let us consider 2 clusters in this case.
Clustering Hamiltonian#
Clustering can be done by minimizing the following Hamiltonian:
is the sample number, is the distance between the two samples, and is a spin variable that indicates which of the two clusters it belongs to. Each term of this Hamiltonian sum is:
0 when
when
With the negative on the right-hand side of the Hamiltonian, the entire Hamiltonian comes down to the question to choose the pair of that maximizes the distance between samples belonging to different classes.
Import Libraries#
We use JijModeling for modeling and JijModeling Transpiler for QUBO generation.
import jijmodeling as jm
import numpy as np
import openjij as oj
import pandas as pd
from jijmodeling.transpiler.pyqubo.to_pyqubo import to_pyqubo
from matplotlib import pyplot as plt
from scipy.spatial import distance_matrix
---------------------------------------------------------------------------
ModuleNotFoundError Traceback (most recent call last)
Cell In[1], line 1
----> 1 import jijmodeling as jm
2 import numpy as np
3 import openjij as oj
ModuleNotFoundError: No module named 'jijmodeling'
Clustering with JijModeling and OpenJij#
First, we formulate the above Hamiltonian using JijModeling. Since JijModeling cannot handle the spin variable , we rewrite it using the relation so that it can be written in the binary variable .
problem = jm.Problem("clustering")
d = jm.Placeholder("d", dim=2)
N = d.shape[0].set_latex("N")
x = jm.Binary("x", shape=(N))
i = jm.Element("i", (0, N))
j = jm.Element("j", (0, N))
problem += (
-1 / 2 * jm.Sum([i, j], d[i, j] * (1 - (2 * x[i] - 1) * (2 * x[j] - 1)))
)
problem
Generating Artificial Data#
Let us artificially generate data that is linearly separable on a 2D plane.
data = []
label = []
for i in range(100):
# Generate random numbers in [0, 1]
p = np.random.uniform(0, 1)
# Class 1 when a condition is satisfied, -1 when it is not.
cls =1 if p>0.5 else -1
# Create random numbers that follow a normal distribution
data.append(np.random.normal(0, 0.5, 2) + np.array([cls, cls]))
label.append(cls)
# Format as a DataFrame
df1 = pd.DataFrame(data, columns=["x", "y"], index=range(len(data)))
df1["label"] = label
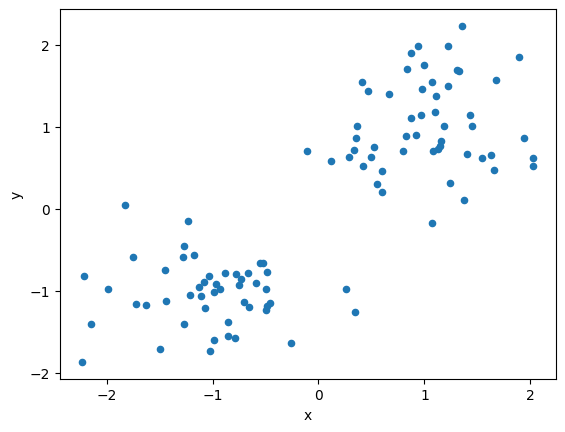
# Visualize dataset
df1.plot(kind='scatter', x="x", y="y")
<AxesSubplot: xlabel='x', ylabel='y'>
instance_data = {"d": distance_matrix(df1, df1)}
Solving the Clustering Problem using OpenJij#
With the mathematical model and data, let us start solving the problem by Openjij. Here we use JijModeling Transpiler.
pyq_obj, pyq_cache = to_pyqubo(problem, instance_data, {})
qubo, constant = pyq_obj.compile().to_qubo()
sampler = oj.SASampler()
response = sampler.sample_qubo(qubo)
result = pyq_cache.decode(response)
# visualize
for idx in range(0, len(instance_data['d'])):
if idx in result.record.solution["x"][0][0][0]:
plt.scatter(df1.loc[idx]["x"], df1.loc[idx]["y"], color="b")
else:
plt.scatter(df1.loc[idx]["x"], df1.loc[idx]["y"], color="r")
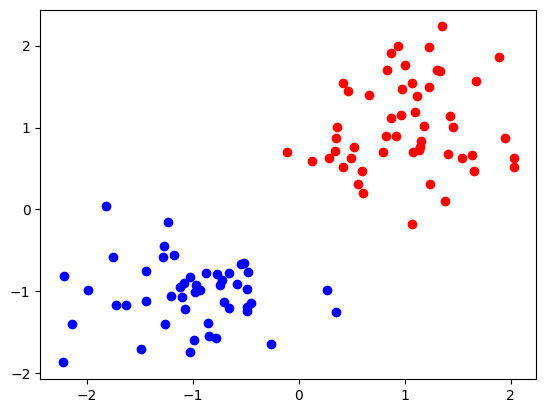
We see that they are classified into red and blue classes.